FVGP MULTI Task Notebook
In this notebook we will go through many features of FVGP. We will be primarily concerned with regression over a single dimension output and multiple tasks.
This first cell has nothing to do with gpCAM, it’s just a function to plot some results later
import plotly.graph_objects as go
import numpy as np
def plot(x,y,z,data = None):
fig = go.Figure()
fig.add_trace(go.Surface(x = x, y = y,z=z))
if data is not None:
fig.add_trace(go.Scatter3d(x=data[:,0], y=data[:,1], z=data[:,2],
mode='markers'))
fig.update_layout(title='Posterior Mean', autosize=True,
width=800, height=800,
margin=dict(l=65, r=50, b=65, t=90))
fig.show()
Import fvgp and relevant libraries
import fvgp
from fvgp import gp, fvgp
import numpy as np
import matplotlib.pyplot as plt
Defining some input data and testing points
def function(x):
data_1 = 100*np.sin(x)+np.cos(x)
data_2 = 5*np.ones(x.shape)
data_3 = 1*np.cos(x/10 + 5)
data_4 = 5*np.sin(x/200)
data_5 = 10*np.cos(x)
return np.column_stack((data_1, data_2, data_3, data_4, data_5))
x_data = np.linspace(-2*np.pi, 10*np.pi,100).reshape(-1,1)
y_data = function(x_data)
x_pred = np.linspace(3*np.pi, 4*np.pi, 100)
Setting up the fvgp multi task object
obj = fvgp.fvGP(1,1,5,x_data,y_data,
init_hyperparameters = np.array([10,10,10]))
Training our gaussian process regression on given data
hyper_param_bounds = np.array([[0.0001, 1000],[ 0.0001, 1000],[ 0.0001, 1000]])
obj.train(hyper_param_bounds)
---------------------------------------------------------------------------
KeyboardInterrupt Traceback (most recent call last)
Cell In[8], line 2
1 hyper_param_bounds = np.array([[0.0001, 1000],[ 0.0001, 1000],[ 0.0001, 1000]])
----> 2 obj.train(hyper_param_bounds)
File ~/checkouts/readthedocs.org/user_builds/fvgp/envs/stable/lib/python3.8/site-packages/fvgp/gp.py:279, in GP.train(self, hyperparameter_bounds, init_hyperparameters, method, pop_size, tolerance, max_iter, local_optimizer, global_optimizer, constraints, deflation_radius, dask_client)
276 if init_hyperparameters is None:
277 init_hyperparameters = np.array(self.hyperparameters)
--> 279 self.hyperparameters = self._optimize_log_likelihood(
280 init_hyperparameters,
281 np.array(hyperparameter_bounds),
282 method,
283 max_iter,
284 pop_size,
285 tolerance,
286 constraints,
287 local_optimizer,
288 global_optimizer,
289 deflation_radius,
290 dask_client
291 )
292 self._compute_prior_fvGP_pdf()
File ~/checkouts/readthedocs.org/user_builds/fvgp/envs/stable/lib/python3.8/site-packages/fvgp/gp.py:476, in GP._optimize_log_likelihood(self, starting_hps, hp_bounds, method, max_iter, pop_size, tolerance, constraints, local_optimizer, global_optimizer, deflation_radius, dask_client)
474 logger.debug("termination tolerance: {}", tolerance)
475 logger.debug("bounds: {}", hp_bounds)
--> 476 res = differential_evolution(
477 self.neg_log_likelihood,
478 hp_bounds,
479 maxiter=max_iter,
480 popsize = pop_size,
481 tol = tolerance,
482 constraints = constraints,
483 workers = 1,
484 )
485 hyperparameters = np.array(res["x"])
486 Eval = self.neg_log_likelihood(hyperparameters)
File ~/checkouts/readthedocs.org/user_builds/fvgp/envs/stable/lib/python3.8/site-packages/scipy/optimize/_differentialevolution.py:397, in differential_evolution(func, bounds, args, strategy, maxiter, popsize, tol, mutation, recombination, seed, callback, disp, polish, init, atol, updating, workers, constraints, x0, integrality, vectorized)
380 # using a context manager means that any created Pool objects are
381 # cleared up.
382 with DifferentialEvolutionSolver(func, bounds, args=args,
383 strategy=strategy,
384 maxiter=maxiter,
(...)
395 integrality=integrality,
396 vectorized=vectorized) as solver:
--> 397 ret = solver.solve()
399 return ret
File ~/checkouts/readthedocs.org/user_builds/fvgp/envs/stable/lib/python3.8/site-packages/scipy/optimize/_differentialevolution.py:998, in DifferentialEvolutionSolver.solve(self)
995 for nit in range(1, self.maxiter + 1):
996 # evolve the population by a generation
997 try:
--> 998 next(self)
999 except StopIteration:
1000 warning_flag = True
File ~/checkouts/readthedocs.org/user_builds/fvgp/envs/stable/lib/python3.8/site-packages/scipy/optimize/_differentialevolution.py:1385, in DifferentialEvolutionSolver.__next__(self)
1383 feasible = True
1384 cv = np.atleast_2d([0.])
-> 1385 energy = self.func(parameters)
1386 self._nfev += 1
1388 # compare trial and population member
File ~/checkouts/readthedocs.org/user_builds/fvgp/envs/stable/lib/python3.8/site-packages/scipy/_lib/_util.py:372, in _FunctionWrapper.__call__(self, x)
371 def __call__(self, x):
--> 372 return self.f(x, *self.args)
File ~/checkouts/readthedocs.org/user_builds/fvgp/envs/stable/lib/python3.8/site-packages/fvgp/gp.py:587, in GP.neg_log_likelihood(self, hyperparameters)
585 mean = self.mean_function(self.x_data,hyperparameters,self)
586 if mean.ndim > 1: raise Exception("Your mean function did not return a 1d numpy array!")
--> 587 x,K = self._compute_covariance_value_product(hyperparameters,self.y_data, self.variances, mean)
588 y = self.y_data - mean
589 sign, logdet = self.slogdet(K)
File ~/checkouts/readthedocs.org/user_builds/fvgp/envs/stable/lib/python3.8/site-packages/fvgp/gp.py:709, in GP._compute_covariance_value_product(self, hyperparameters, values, variances, mean)
708 def _compute_covariance_value_product(self, hyperparameters,values, variances, mean):
--> 709 K = self._compute_covariance(hyperparameters, variances)
710 y = values - mean
711 x = self.solve(K, y)
File ~/checkouts/readthedocs.org/user_builds/fvgp/envs/stable/lib/python3.8/site-packages/fvgp/gp.py:717, in GP._compute_covariance(self, hyperparameters, variances)
715 def _compute_covariance(self, hyperparameters, variances):
716 """computes the covariance matrix from the kernel"""
--> 717 CoVariance = self.kernel(
718 self.x_data, self.x_data, hyperparameters, self)
719 self.add_to_diag(CoVariance, variances)
720 return CoVariance
File ~/checkouts/readthedocs.org/user_builds/fvgp/envs/stable/lib/python3.8/site-packages/fvgp/gp.py:1718, in GP.default_kernel(self, x1, x2, hyperparameters, obj)
1716 distance_matrix += abs(np.subtract.outer(x1[:,i],x2[:,i])/hps[1+i])**2
1717 distance_matrix = np.sqrt(distance_matrix)
-> 1718 return hps[0] * obj.matern_kernel_diff1(distance_matrix,1)
File ~/checkouts/readthedocs.org/user_builds/fvgp/envs/stable/lib/python3.8/site-packages/fvgp/gp.py:1492, in GP.matern_kernel_diff1(self, distance, length)
1473 def matern_kernel_diff1(self, distance, length):
1474 """
1475 Function for the matern kernel, order of differentiablity = 1.
1476 kernel = (1.0 + ((np.sqrt(3.0) * distance) / (length))) * np.exp(
(...)
1488 A structure of the shape of the distance input parameter : float or np.ndarray
1489 """
1491 kernel = (1.0 + ((np.sqrt(3.0) * distance) / (length))) * np.exp(
-> 1492 -(np.sqrt(3.0) * distance) / length
1493 )
1494 return kernel
KeyboardInterrupt:
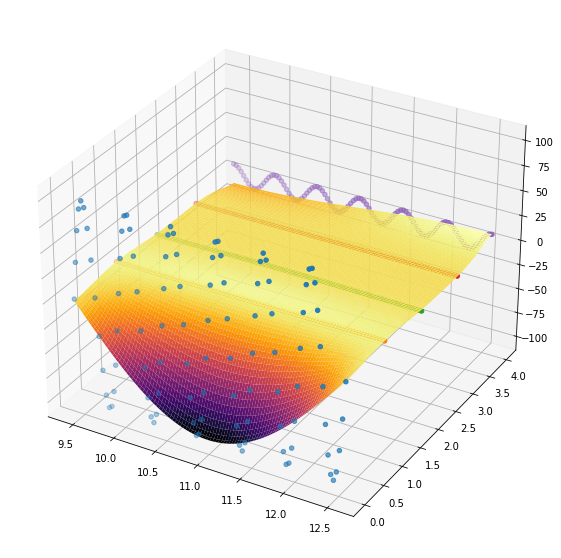
Looking at the posterior mean at the test points (remember that we did not define a particularly good kernel)
task_idx = 1
x_linspace = np.linspace(3*np.pi, 4*np.pi,100)
y_linspace = np.linspace(0,4,100)
x_grid, y_grid = np.meshgrid(x_linspace, y_linspace)
post_mean= obj.posterior_mean(np.column_stack((x_pred, task_idx*np.ones(x_pred.shape))))
posterior_mean = obj.posterior_mean(np.column_stack((x_grid.flatten(), y_grid.flatten())))
fig = plt.figure(figsize = (10,10))
ax = fig.add_subplot(projection='3d')
ax.scatter(post_mean['x'][:,0], np.ones(100)*0,y_data[:,0])
ax.scatter(post_mean['x'][:,0], np.ones(100)*1,y_data[:,1])
ax.scatter(post_mean['x'][:,0], np.ones(100)*2,y_data[:,2])
ax.scatter(post_mean['x'][:,0], np.ones(100)*3,y_data[:,3])
ax.scatter(post_mean['x'][:,0], np.ones(100)*4,y_data[:,4])
ax.plot_surface(x_grid, y_grid, posterior_mean['f(x)'].reshape(100,100), rstride=1, cstride=1,cmap='inferno')
<mpl_toolkits.mplot3d.art3d.Poly3DCollection at 0x7f3a2804daf0>